ABC_GWH
A cookbook to study Genome Wide Heterogeneity in introgression rates
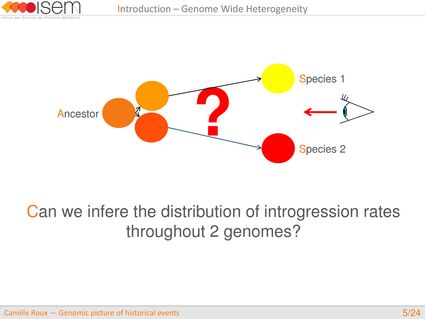
How many loci cross the species barrier because of adaptive introgression?
How are they distributed throughout genomes?
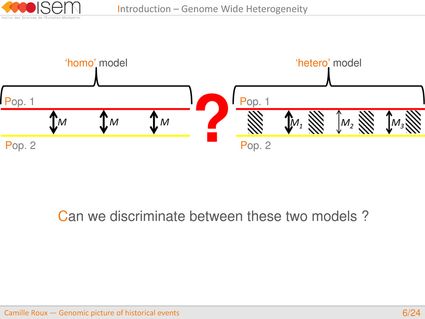
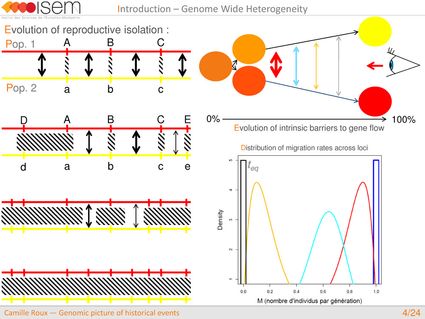
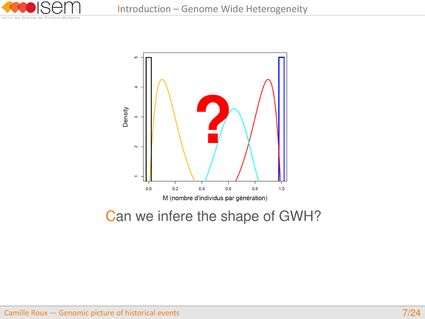
What is the global shape of the genome-wide distribution in introgression rates between two gene pools (GWH)?
Here we propose a cookbook to approach these questions from datasets describing patterns of molecular divergence and polymorphism.
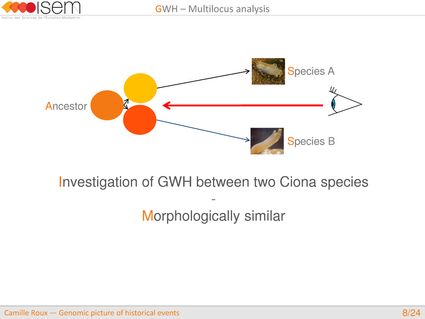
To illustrate how these issues can be addressed empirically, we now present our study on speciation between Ciona intestinalis A and C. intestinalis B. Even if these two species are morphologically very similar, they are highly diverged molecular-wise with a synonymous divergence of 14.4%, i.e, equivalent to the divergence between humans and marmosets.


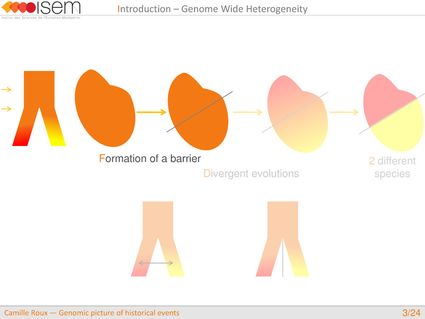
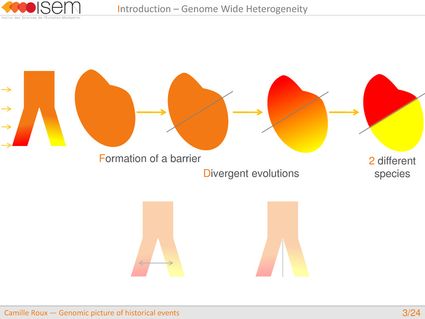
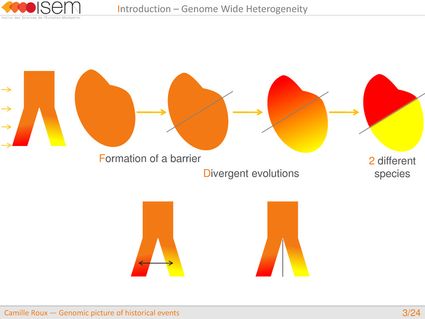
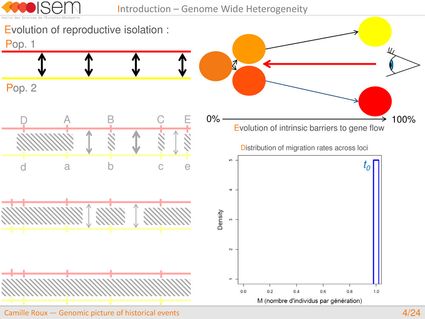
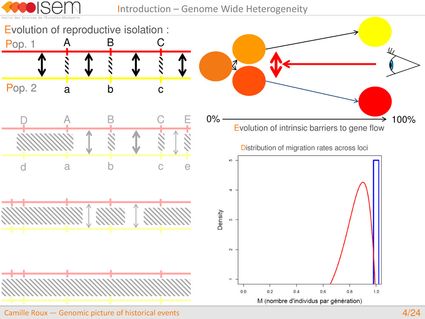
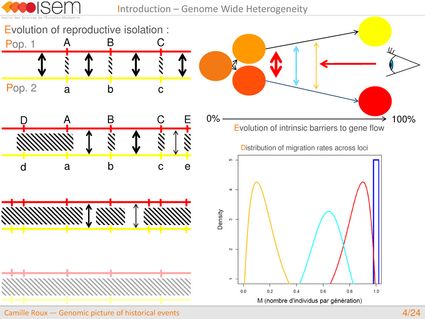

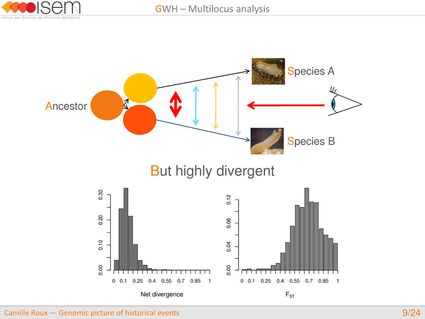
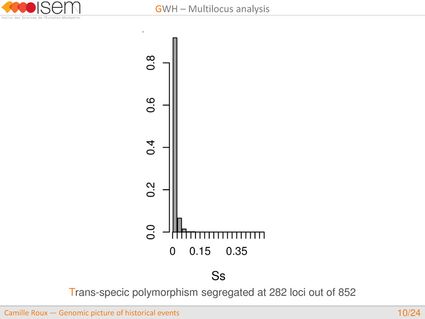
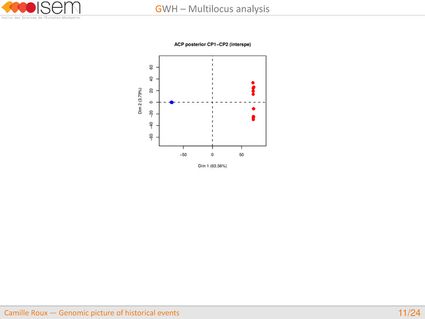
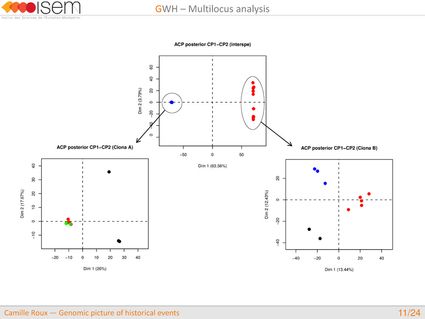
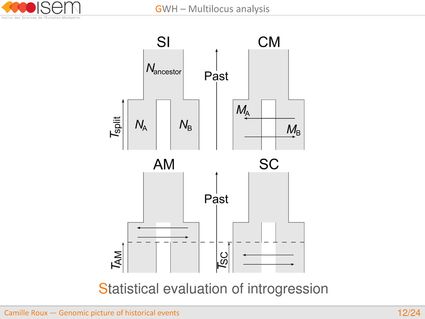


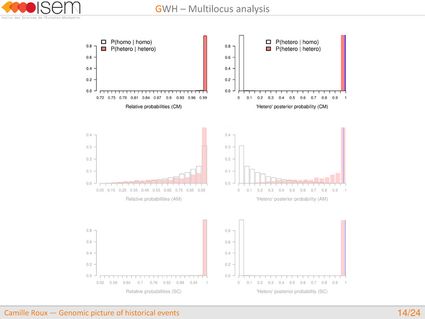
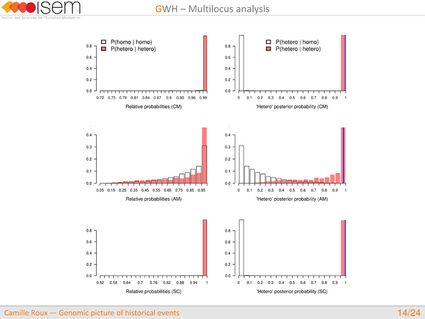
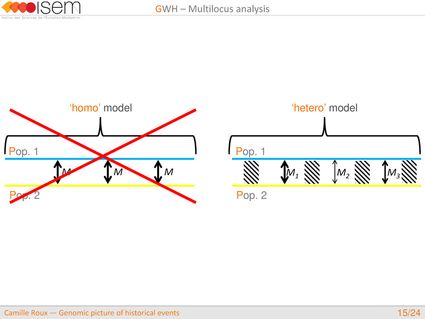
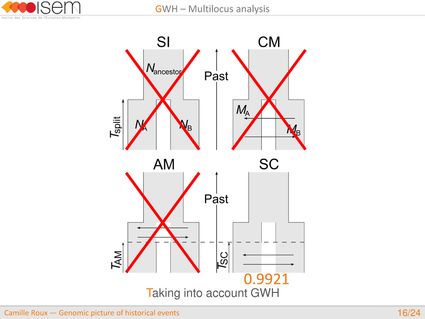
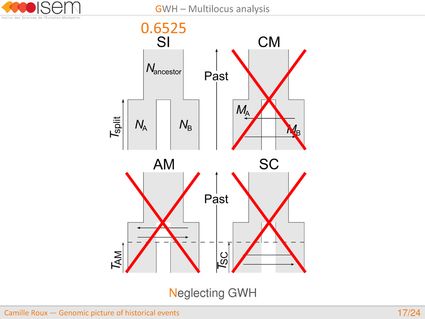
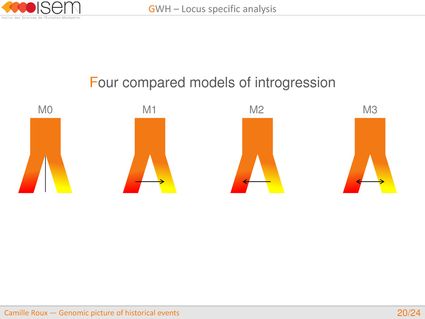
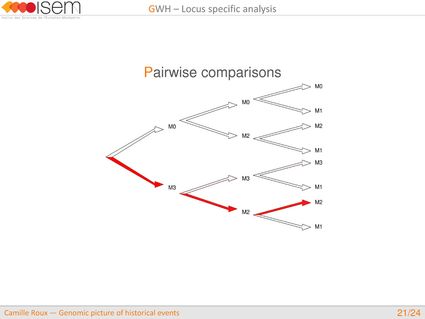
In spite of this long history of divergence, evidence of hybridisation has been reported both in the laboratory and in field studies. We investigated genome-wide heterogeneity (GWH) of the rate of introgression as a genomic outcome of the speciation process, but also asked how this process could affect statistical inference of divergence with gene flow.
The main results from this study are :
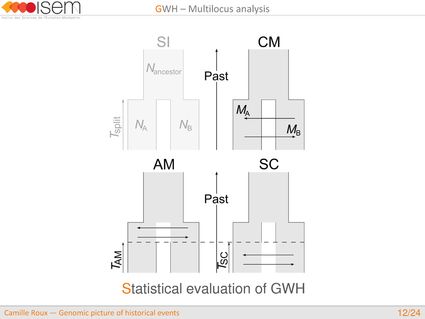
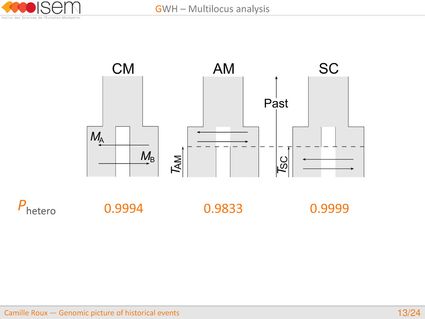
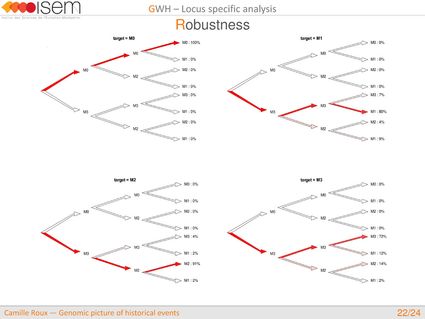
1. Testing for GWH of the rate of introgression is highly robust in an ABC framework.
2. Neglecting the effects of GWH may lead to wrong biological conclusions, while GWH is more likely the general rule than the exception in nature.
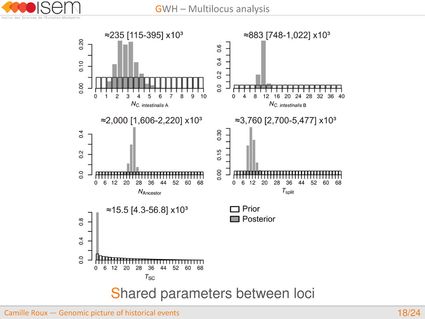
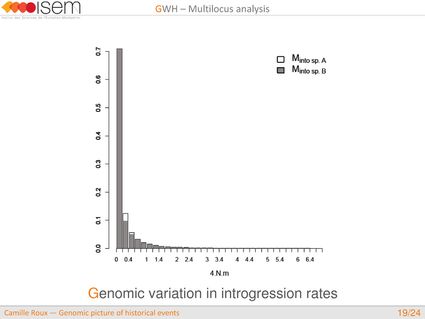
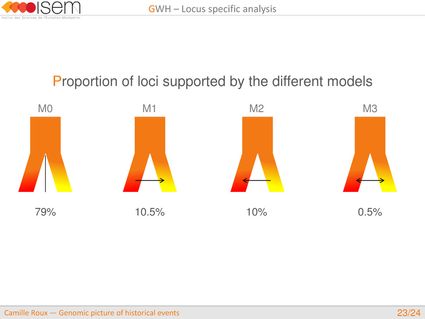
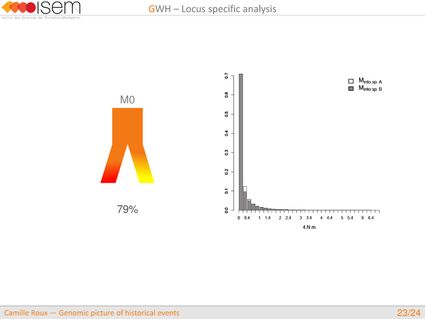
3. Between the two Ciona species, GWH of the rate of introgression is leptokurtic (L-shaped), as expected in the final stages of evolutionary divergence, after the build-up of numerous barriers to gene flow throughout the genome.
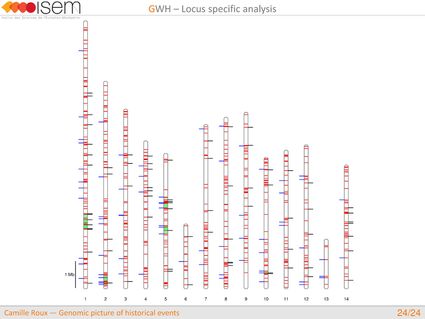
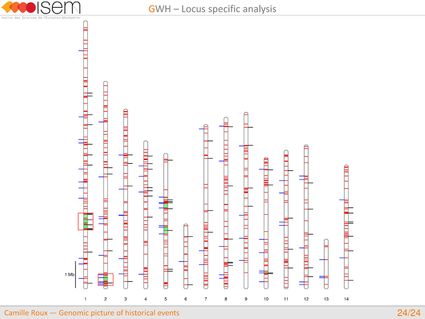
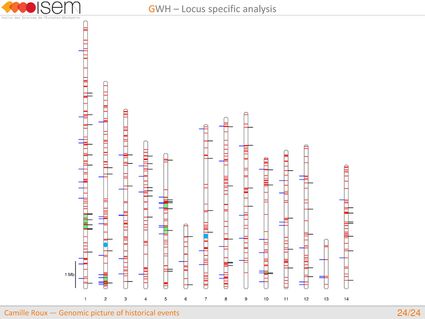
4. We detected genomic islands of introgression, i.e, genomic regions with a high concentration of loci at which the two species are particularly prone to exchange alleles.